SS320 (MC1061 F') Electrocompetent Cells
Product Details
ShowCompetent Cells include Control DNA and Recovery Medium and are packaged as DUOs (2 transformations per tube). Recovery Medium is also available separately. The specified transformation efficiencies are with pUC DNA, unless indicated otherwise.
SS320 (MC1061 F') Electrocompetent Cells
Key features
Show- Perfect for antibody phage display library creation
- Create larger libraries; speed discovery
- Bulk custom dispensing available with all of our phage display cells
Product information
SS320 Electrocompetent Cells. A non-amber suppressor strain (sometimes called MC1061F') prepared as highly efficient (≥4 × 1010 cfu/µg) electrocompetent cells for phage display library screening. These cells have the highest available transformation efficiency of any strain used for phage display.
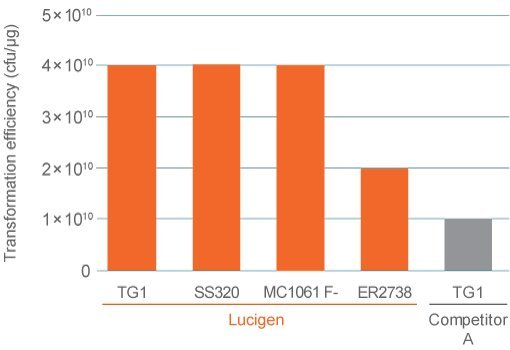
Figure 1. Transformation efficiency of electrocompetent bacterial cells for phage display compared to competitor's specification.
Table 1. Competent Cells for Phage Display libraries
Cell Lines |
Transformation Efficiency |
Cloning Methylated DNA |
BAC, Cosmid Cloning |
Blue/White Screening |
|
TG1 Electrocompetent |
>4 × 1010 |
YES |
NO |
YES |
|
SS320 (MC1061 F') Electrocompetent |
>4× 1010 |
YES |
NO |
YES |
|
MC1061 F-Electrocompetent |
>4× 1010 |
YES |
NO |
YES |
|
ER2738 Electrocompetent |
>2 × 1010 |
YES |
NO |
YES |
Genotype
- TG1: [F' traD36 proAB lacIqZ ΔM15] supE thi-1 Δ(lac-proAB) Δ(mcrB-hsdSM)5(rK - mK -)
- SS320 (MC1061F'): F'[proAB lacIqZ ΔM15 Tn10 (TetR)] araD139 Δ(ara-leu)7696 galE15 galK16 Δ(lac)X74 rpsL (StrR) hsdR2 (rK– mK+) mcrA mcrB1
- ER2738: [F'proA+B+ lacIq Δ(lacZ)M15 zzf::Tn10 (tetr)] fhuA2 glnVΔ(lac-proAB) thi-1Δ(hsdS-mcrB)5
- MC1061 F-: F-, araD139 Δ(ara-leu)7696 galE15 galK16 Δ(lac)X74 rpsL (StrR) hsdR2 (rK– mK+) mcrA mcrB1
SDS
-
SDS007 60512 SS320 MC1061 F Prime Electrocompetent Cells
-
60512 SS320 MC1061 F Prime Electrocompetent Cells
Manuals and user guides
Product information sheets
Access support
Need some support with placing an order, setting up an account, or finding the right protocol?
Contact us